stormchasegenie
Member
Greetings,
I am downscaling a very low-resolution GCM (MPI-ESM1-2-LR) using WRF. I am using skin temperature (SKINTEMP) from the GCM output to generature the intermediate binaries.
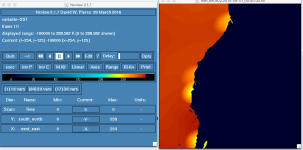
When generated, the me_em files show blocky patterns in the SST field along the shoreline which precisely match the LANDSEA mask. Is there a standard way to smooth out these artifacts using metgrid.exe?
All my best, and thank you.
-Stefan
I am downscaling a very low-resolution GCM (MPI-ESM1-2-LR) using WRF. I am using skin temperature (SKINTEMP) from the GCM output to generature the intermediate binaries.
When generated, the me_em files show blocky patterns in the SST field along the shoreline which precisely match the LANDSEA mask. Is there a standard way to smooth out these artifacts using metgrid.exe?
All my best, and thank you.
-Stefan